
GIPZ
™
Lentiviral shRNA
TECHNICAL MANUAL
horizondiscovery.com
Product description
The Dharmacon
™
GIPZ
™
Lentiviral shRNA Library was developed in collabora-
tion with Dr. Greg Hannon of Cold Spring Harbor Laboratory [CSHL] and Dr.
Steve Elledge of Harvard Medical School. This library combines the design
advantages of microRNA-adapted shRNA with the pGIPZ lentiviral vector to
create a powerful RNA tool capable of producing RNA interference (RNAi) in
most cell types including primary and non-dividing cells.
Important safety note
Please follow the safety guidelines for use and production of vector-based
lentivirus as set by your institution’s biosafety committee.
• For glycerol stocks of E. coli containing lentiviral plasmids, BSL1 guidelines
should be followed
• For handling and use of lentiviral products to produce lentiviral particles,
BSL2 or BSL2+ guidelines should be followed
• For handling and use of lentiviral particle products, BSL2 or BSL2+ guide-
lines should be followed
Additional information on the safety features incorporated in the pGIPZ
lentiviral vector and the Horizon Trans-Lentiviral Packaging System can be
found on page 3.
Design information
Unique microRNA-30 based hairpin design
Short hairpin RNA (shRNA) constructs are expressed as human microRNA-30
(miR-30) primary transcripts. This design adds a Drosha processing site to the
hairpin construct and has been shown to greatly increase gene silencing
eciency (Boden 2004). The hairpin stem consists of 22 nucleotides (nt) of
dsRNA and a 19 nucleotides (nt) loop from human miR-30. Adding the miR-30
loop and 125 nucleotides (nt) of miR-30 anking sequence on either side of the
hairpin results in greater than 10-fold increase in Drosha and Dicer processing
of the expressed hairpins when compared with conventional shRNA designs
(Silva 2005). Increased Drosha and Dicer processing translates into greater
shRNA production and greater potency for expressed hairpins.
Use of the miR-30 design also allowed the use of ‘rules-based’ designs for target
sequence selection. One such rule is the destabilizing of the 5' end of the
antisense strand, which results in strand specic incorporation of microRNA/
siRNAs into RISC. The proprietary design algorithm targets sequences in coding
regions and the 3' UTR with the additional requirement that they contain great-
er than 3 mismatches to any other sequence in the human or mouse genomes.
Each shRNA construct has been bioinformatically veried to match NCBI
sequence data. To assure the highest possibility of modulating the gene
expression level, each gene is represented by multiple shRNA constructs, each
covering a unique region of the target gene.
Vector information
Versatile vector design
Features of the pGIPZ lentiviral vector (Figures 1 and 2) that make it a versatile
tool for RNAi studies include:
• Ability to perform transfections or transductions using the replication
incompetent lentivirus (Shimada 1995)
• TurboGFP™ (Evrogen, Moscow, Russia) and shRNA are part of a bicistronic
transcript allowing the visual marking of shRNA expressing cells.
• Amenable to in vitro and in vivo applications.
• Puromycin drug resistance marker for selecting stable cell lines.
• Molecular barcodes enable multiplexed screening in pools
Dharmacon
™
GIPZ
™
shRNA vectors are not compatible with
third generation packaging systems, due to the requirement
of the expression of tat, which third generation systems do
not contain. We recommend the Trans-Lentiviral Packaging
System for use with our vectors.
GIPZ shRNA is available in glycerol stock or viral particle
format. If viral particle format is purchased, begin work with
Protocol IX – Determining Relative Transduction Efficiency.

pGIPZ
11.8 kb
Drawing was created using Geneious (http://www.geneious.com).
Vector map
Figure 2. Detailed vector map of pGIPZ lentiviral vector.
Antibiotic resistance
pGIPZ contains three antibiotic resistance markers (Table 2).
Puro
R
IRES
Amp
R
SV40 ori
pUC ori
tGFP
hCMV
RRE
shRNA
WPRE
5' LTR
3' SIN LTR
Ψ
pGIPZ
pTRIPZ
rtTA3
tRFP
TRE
UBC
From PowerPoint (R&D)Revised versions
pLOC
pLKO.1
U6
hPGK
pSMART 2.0
SV40 ori
hCMV
pSMART 2.0
pSMART 2.0
Amp
R
pUCori
SV40 ori
5ʹ LTR
3ʹ SIN LTR
Ψ
RRE
tGFP
hCMV
IRES
Puro
R
shRNA WPRE
cPPT/CTS
hPGK
U6
Amp
R
pUCori
pLKO.1
RSV/5ʹ LTR
3ʹ SIN LTR
Ψ
RRE
Puro
R
shRNA
pSMART 2.0
Amp
R
pUCori
SV40 ori
5ʹ LTR
3ʹ SIN LTR
Ψ
RRE
tGFP
hCMV
IRES
Puro
R
microRNA
WPRE
pLOC
Amp
R
pUCori
SV40 ori
5ʹ LTR
3ʹ SIN LTR
Ψ
RRE
tGFP
nuc
hCMV
IRES
Blast
R
-2a-
WPRE
ORF
Multi Tag
Cloning Site
Amp
R
pUCori
SV40 ori
pGIPZ
5ʹ LTR
3ʹ SIN LTR
Ψ
RRE
tGFP
hCMV
IRES
Puro
R
WPRE
shRNA
Amp
R
pUCori
SV40 ori
UBC
pTRIPZ
5ʹ LTR
3ʹ SIN LTR
Ψ
RRE
tRFP
TRE
IRES
Puro
R
WPRE
rtTA3
shRNA
hCMV
mCMV
hEF1α
mEF1α
CAG
PGK
UBC
SMARTchoiceshRNA
5ʹ LTR
3ʹ SIN LTR
Ψ
RRE
tGFP
or
tRFP
IRES
Puro
R
WPRE
SMARTc hoice
promoters
SMARTvector2.0 universal scaold
5' LTR
Ψ
Puro
R
IRES
WPRE
RRE
shRNA
3' SIN LTR
Amp
R
SV40 ori
pUC ori
hCMV
5' LTR
Ψ
5' LTR
Ψ
RSV/5' LTR
Ψ
Puro
R
Puro
R
IRES
WPRE
RRE
RRE
RRE
shRNA
shRNA
3' SIN LTR
3' SIN LTR
3' SIN LTR
5' LTR
Ψ
3' SIN LTR
Amp
R
Amp
R
Amp
R
SV40 ori
pUC ori
pUC ori
pUC ori
SV40 ori
Amp
R
pUC ori
tGFP
Puro
R
IRES
WPRE
microRNA
tGFP
hCMV
Blast
R
IRES
WPRE
-2a-
tGFP
nuc
Multi Tag
Cloning Site
ORF
hCMV
mCMV
hEF1
or
α
mEF1α
CAG
PGK
UBC
SMARTchoice
promoters
5' LTR
Ψ
tGFP
Puro
R
IRES
tRFP
WPRE
3' SIN LTR
SMARTvector universal scaold
SMARTchoice shRNA
RRE
Vector element Utility
hCMV Human cytomegalovirus promoter drives strong transgene
expression
tGFP TurboGFP reporter for visual tracking of transduction and
expression
Puro
R
Puromycin resistance permits antibiotic-selective pressure
and propagation of stable integrants
IRES Internal ribosomal entry site allows expression of TurboGFP
and puromycin resistance genes in a single transcript
shRNA microRNA-adapted shRNA (based on miR-30) for
gene knockdown
5' LTR 5' long terminal repeat
3' SIN LTR 3' self-inactivating long terminal repeat for increased
lentivirus safety
Ψ
Psi packaging sequence allows viral genome packaging
using lentiviral packaging systems
RRE Rev response element enhances titer by increasing
packaging efficiency of full-length viral genomes
WPRE Woodchuck hepatitis posttranscriptional regulatory
element enhances transgene expression in the target cells
Table 1. Features of the pGIPZ vector.
Figure 1. pGIPZ lentiviral vector.
Antibiotic Concentration Utility
Ampicillin (carbenicillin) 100 μg/mL Bacterial selection marker
(outside LTRs)
Zeocin
™
25 μg/mL Bacterial selection marker
(inside LTRs)
Puromycin Variable Mammalian selection marker
Table 2. Antibiotic resistances conveyed by pGIPZ.
Quality control
The GIPZ Lentiviral shRNA Library has passed through internal QC processes to
ensure high quality and low recombination (Figures 3 and 4).
Figure 3. Representative GIPZ Lentiviral shRNA clones grown for 16 hours at 30 °C.
Plasmid was isolated and normalized to a standard concentration. Clones were then
digested with SacII and run on an agarose gel with uncut plasmid. The expected band
sizes are 1259 bp, 2502 bp, and 7927 bp. No recombinant products are visible. 10 kb
molecular weight ladder (10 kb, 7 kb, 5 kb, 4 kb, 3 kb, 2.5 kb, 2 kb, 1.5 kb, 1 kb).
Figure 4. Gel image of a restriction digest of clones from the GIPZ shRNA library
cultured for 10 successive generations in an attempt to determine the tendency of the
pGIPZ vector to recombine. Each generation was thawed, replicated and incubated
overnight for 16 hours at 30 °C then frozen, thawed and replicated. This process was
repeated for 10 growth cycles. After the 10th growth cycle, plasmid was isolated and
normalized to a standard concentration. Clones were digested with SacII and run on
an agarose gel. The pGIPZ vector appears stable without showing any recombination.
Expected band sizes: 1259 bp, 2502 bp, and 7927 bp. 10 kb molecular weight ladder
(10 kb, 7 kb, 5 kb, 4 kb, 3 kb, 2.5 kb, 2 kb, 1.5 kb, 1 kb).
Additional safety information
Historically, the greatest safety risk associated with a lentiviral delivery
platform stems from the potential generation of recombinant viruses that
are capable of autonomous replication. The pGIPZ Lentiviral shRNA platform
minimizes these hazards to the greatest degree by combining a disabled viral
genome with the proprietary Trans-Lentiviral packaging process. Starting with
the HXB2 clone of HIV-1 (GenBank, Accession #K03455), the lentiviral backbone
has been modied to eliminate all but the most essential genetic elements
necessary for packaging and integration (such as 5' LTR, Psi sequences, polypu-
rine tracts, Rev responsive elements and 3' LTR). The resultant self-inactivating
(SIN) vector greatly reduces the probability of producing recombinant particles
and limits cellular toxicity often associated with expression of HIV genes.
horizondiscovery.com

Additional safety features can be incorporated by the packaging process
itself. Generation of pGIPZ Lentiviral shRNA particles requires a packaging
step during which the expression construct containing the silencing sequence
is enclosed in a viral capsid. Gene functions that facilitate this process (such
as those encoded by the structural genes gag, pol, env, etc.) are distributed
amongst multiple helper plasmids which do not contain signicant regions
of homology. This tactic further minimizes the probability of recombination
events that might otherwise generate viruses capable of autonomous
replication. Among commercially available lentiviral vector systems, the
Trans-Lentiviral Packaging System oers a superior safety prole as the
packaging components are separated onto ve plasmids. Additionally,
expression of gag-pro and tat-rev are under the control of the conditional
tetracycline-responsive promoter element (TRE), limiting expression of
these viral components strictly to the packaging cell line. A detailed description
of the Trans-Lentiviral Packaging System can be found in (Wu 2000).
With these safety measures in place, GIPZ lentiviral shRNA particles can be
employed in standard Biosafety Level 2 tissue culture facilities.
Any investigator who purchases Horizon viral vector products is responsible
for consulting with their institution’s health and biosafety group for specic
guidelines on the handling of lentiviral vector particles. Further, each
investigator is fully responsible for obtaining the required permissions for the
acceptance of lentiviral particles into their local geography and institution.
• In the U.S., download the U.S. Department of Health and Human Services
Centers for Disease Control and Prevention and National Institutes of
Health, Biosafety in Microbiological and Biomedical Laboratories (BMBL),
Fifth Edition, Feb 2007 here.
• See also: NIH Guidelines For Research Involving Recombinant DNA
Molecules (NIH Guidelines), September 2009, downloadable here.
• For Biosafety Considerations for Research with Lentiviral Vectors, see.
Replication of individual clones
Once the clone has been streak isolated and the identity of the strain has been
conrmed** by Sanger sequencing (See: What is the sequencing primer for
pGIPZ?), we recommend making a stock of the pure culture. Grow the pure
culture in LB broth + appropriate antibiotic (See protocol below: Protocol
1 - replication). Vortex the culture to evenly mix the glycerol throughout the
culture. The culture can be stored indenitely at –80 °C.
**Testing of 3-5 colonies is recommended.
Protocol I – replication
For archive replication, grow GIPZ shRNA clones at 30 °C in 2x LB broth
(low salt)* medium plus 25 μg/mL Zeocin
™
and 100 μg/mL carbenicillin in order
to provide maximum stability of the clones. Prepare medium with 8% glycerol**
and the appropriate antibiotics.
2x LB broth (low-salt) medium preparation
LB-Broth-Lennox 20 g/L
Peptone 10 g/L
Yeast Extract 5 g/L
Appropriate antibiotic(s) at recommended concentration(s). Glycerol 8% for
long-term storage.
*1x LB medium can be used instead of 2x LB broth medium.
**Glycerol can be omitted from the medium if you are culturing for plasmid
preparation. If making copies of the constructs for long-term storage at
–80 °C, 8% glycerol is required
Table 3. Materials for plate replication.
Item Vendor Cat #
2x LB Broth (low salt) Fisher Scientific BP1427500
Peptone, granulated, 2 kg – Difco Fisher Scientific BP9725-2
Yeast Extract, 500 g, granulated Fisher Scientific BP1422-500
NaCl Fisher Scientific BP3581
Glycerol Fisher Scientific BP2291
Carbenicillin Fisher Scientific BP2648-250
Zeocin Invitrogen ant-zn-5p
Puromycin Fisher Scientific BP2956-100
96-well microplates Fisher Scientific 12-565-363
Aluminum seals Fisher Scientific 12-565-475
Disposable replicators Fisher Scientific NC9584102
Replication of plates
Prepare target plates by dispensing ~ 160 μL of 2x LB broth (low salt) medium
supplemented with 8% glycerol** and appropriate antibiotic (25 μg/mL Zeocin
and 100 μg/mL carbenicillin).
Prepare source plates
Remove foil seals from the source plates while they are still frozen. This
minimizes cross-contamination. Thaw the source plates with the lid on. Wipe any
condensation underneath the lid with a paper wipe soaked in ethanol.
Replicate
1. Gently place a disposable replicator in the thawed source plate and lightly
move the replicator around inside the well to mix the culture. Make sure to
scrape the bottom of the plate of the well.
2. Gently remove the replicator from the source plate and gently place in the
target plate and mix in the same manner to transfer cells.
3. Dispose of the replicator.
4. Place the lids back on the source plates and target plates.
5. Repeat steps 1-4 until all plates have been replicated.
6. Return the source plates to the -80 °C freezer.
7. Place the inoculated target plates in a 30 °C incubator for 18-19 hours.
Freeze at -80 °C for long-term storage. Avoid long periods of storage at room
temperature or higher in order to control background recombination products.
Protocol II – Plasmid preparation
Culture conditions for individual plasmid preparations
For plasmid preparation, grow all GIPZ shRNA clones at 37 °C in 2x LB broth
(low salt) medium plus 100 μg/mL carbenicillin only.
1. Upon receiving your glycerol stock(s) containing the shRNA of interest,
conrm the clone identity (See: Replication of individual clones) and store
immediately at –80 °C until ready to begin.
2. To prepare plasmid DNA, rst thaw your glycerol stock culture and pulse
vortex to resuspend any E. coli that may have settled to the bottom of
the tube.
3. Take a 10 μL inoculum from the glycerol stock into 3–5 mL of 2x LB broth
(low salt) medium with 100 μg/mL carbenicillin. Return the glycerol stock(s)
to –80 °C.
Dilute the starter culture 1:500–1:1000 into the larger volume.
4. Incubate at 37 °C for 18–19 hours with vigorous shaking.
Due to the tendency of viral vectors to recombine, we
recommend keeping the incubation times as short as
possible and avoid subculturing. Return to your glycerol
stock of your pure culture (see Replication of individual
clones) for each plasmid preparation.
If a large culture volume is desired, incubate the 3-5 mL culture
for 8 hours at 37 ˚C with shaking and use as a starter inoculum.
horizondiscovery.com

Protocol IV – Transfection
Quantities and volumes should be scaled-up according to the number of
cells/wells to be transfected (Table 5). This example is for 24-well plate format.
1. In each well, seed ~ 5 × 10
4
adherent cells or ~ 5 × 10
5
suspension cells in
0.5 mL of growth medium 24 hours prior to transfection.
For the library collection, we use the above 96-well bio-
block plasmid preparation protocol in conjunction with
a Qiagen
™
Turbo
™
Kit (Cat #27191). We use 2 bio-blocks
combined. Do not perform the optional wash and elute the
DNA in molecular grade water.
5. Pellet the culture and begin preparation of plasmid DNA. Plasmid DNA
can be isolated using Thermo Scientic
™
GeneJET
™
Plasmid Miniprep Kit
(Cat #K0502) or similar.
6. Run 0.2–1 μg of the plasmid DNA on a 1% agarose gel. pGIPZ with shRNA
is 11774 bp.
Culture conditions for 96-well bio-block plasmid preparation
Inoculate a 96-well bio-block containing 1 mL per well of 2x LB broth (low
salt) medium with 100 μg/mL carbenicillin with 1 µL of the glycerol stock
culture. Incubate at 37 °C with shaking (~ 170–200 rpm). We have observed
that incubation times between 18–19 hours produce good plasmid yield.
For plasmid preparation, follow the protocols recommended by the plasmid
isolation kit manufacturer.
Due to the tendency of viral vectors to recombine, we
recommend keeping the incubation times as short as
possible and avoid subculturing. Return to your original
glycerol stock of your pure culture (see Replication of
individual clones) for each plasmid preparation.
The recommended confluency for adherent cells on the
day of transfection is 70-90%. Suspension cells should be
in logarithmic growth phase at the time of transfection.
2. Dilute 1 µg of DNA in 50 µL of DMEM or other serum-free growth medium.
3. Gently mix DharmaFECT kb transfection reagent and add 3 µL to the diluted
DNA. Mix immediately by pipetting.
4. Incubate 10 minutes at room temperature. Remove medium from
wells and replace with 0.45 mL fresh growth medium.
5. Gently add 50 µL of the DharmaFECT kb reagent/DNA mixture to each well.
6. Gently rock the plate to achieve even distribution of the complexes.
7. Incubate at 37 °C in a CO
2
incubator.
8. Analyze transgene expression 24-48 hours later. For stable transfection,
cells should be grown in selective medium for 10-15 days (see Protocol V –
Puromycin Selection).
Component Amount
Water, nuclease-free (Cat #R0581) X μL
10x FastDigest
™
buffer 2 μL
DNA sample (up to 1 µg) in water X μL
FastDigest enzyme 1 μL
Final Volume 20 μL
Table 4. Restriction digest components.
1 2 3
4
5 6
Lane 1 - 10 kb molecular weight ladder (10 kb, 7 kb, 5 kb, 4 kb, 3 kb, 2.5 kb,
2 kb, 1.5 kb, 1 kb)
Lane 2 - Uncut pGIPZ vector
Lane 3 - KpnI digested pGIPZ produces two bands at 1917 bp and 9857 bp.
Lane 4 - SacII digest produces three bands at 1345 bp, 2502 bp and 7927 bp
Lane 5 - SalI produces three bands at 2188 bp, 4465 bp and 5121 bp
Lane 6 - XhoI, NotI double digest produces two bands at 1291 bp & 10,483 bp
horizondiscovery.com
Protocol III – restriction digest
The following is a sample protocol for restriction enzyme digestion using
Thermo Scientic
™
FastDigest
™
Restriction Enzymes KpnI (Cat #FD0524), SalI (Cat
#FD0644), XhoI (Cat #FD0694) and/or NotI (Cat #FD0594) and Thermo Scientic
™
Restriction Enzyme
™
SalI (Cat #ER0205) for diagnostic quality control of pGIPZ
lentiviral vectors.
1. Add the following components (Table 4), in the order stated, to a sterile
PCR thin-wall tube.
2. Mix gently by pipetting.
3. Incubate in a thermal cycler at 37 °C for 5 minutes for FastDigest enzymes
or as suggested by the manufacturer.
4. Load the gel with 10 µL of each of the digested samples, KpnI, SacII, SalI,
XhoI and/or NotI on a 1% agarose gel. Run uncut sample alongside the
digested samples. (Figure 5).
Prepare immediately prior to transfection. We recommend
starting with 1 µg of DNA and 3 µL of DharmaFECT kb
reagent per well in a 24-well plate (see scale-up Table 5).
Subsequent optimization may further increase transfection
efficiency depending on the cell line and transgene used.
The transfection efficiency with DharmaFECT kb
transfection reagent (Horizon, Cat #T-2006-01) is equally
high in the presence of serum. This is not the case with
other transfection reagents.
Tissue culture vessel
Growth area,
cm
2
/well
Volume of
medium, mL
Adherent (suspension) cells to seed
the day before transfection*
Amount of DNA Volume of DharmaFECT kb, μL
µg** µL*** Recommended Range
96-well plate 0.3 0.1 0.5-1.2 × 10
4
(2.0 × 10
4
) 0.2 10 0.6 0.4-1.0
48-well plate 0.7 0.25 1.0-3.0 × 10
4
(5.0 × 10
4
) 0.5 25 1.5 0.8-2.2
24-well plate 2.0 0.5 2.0-6.0 × 10
4
(1.0 × 10
5
) 1.0 50 3.0 2.0-5.0
12-well plate 4.0 1.0 0.4-1.2 × 10
5
(2.0 × 10
5
) 2.0 100 6.0 3.9-9.0
6-well plate 9.5 2.0 0.8-2.4 × 10
5
(4.0 × 10
5
) 4.0 200 9.0 6.0-12.0
60 mm plate 20 3.0 2.0-6.3 × 10
5
(1.0 × 10
6
) 6.0 300 18.0 12.0-24.0
* These numbers were determined using HEK293T and U2OS cells. Actual values depend on the cell type.
** Amount of DNA and DharmaFECT kb transfection reagent used may require optimization.
*** The volume of DNA should be 1/10 of the volume of the culture medium used for dilution of the DNA.
Table 3. Scale-up ratios for transfection of adherent and suspensioncells with DharmaFECT kb transfection reagent
Protocol VII – Titering
Viral titering
Follow the procedure below to determine the titer of your lentiviral stock
using the mammalian cell line of choice IF YOU HAVE PRODUCED VIRAL
PARTICLES YOURSELF. This protocol uses the HEK293T cell line that is
available as part of our Trans-Lentiviral shRNA Packaging Kit (Cat #TLP5918).
You can use a standard HEK293T cell line as an alternative.
1. The day before transduction, seed a 24-well tissue culture plate
with HEK293T Cells at 5 × 10
4
cells per well in DMEM (10% FBS,
1% pen-strep).
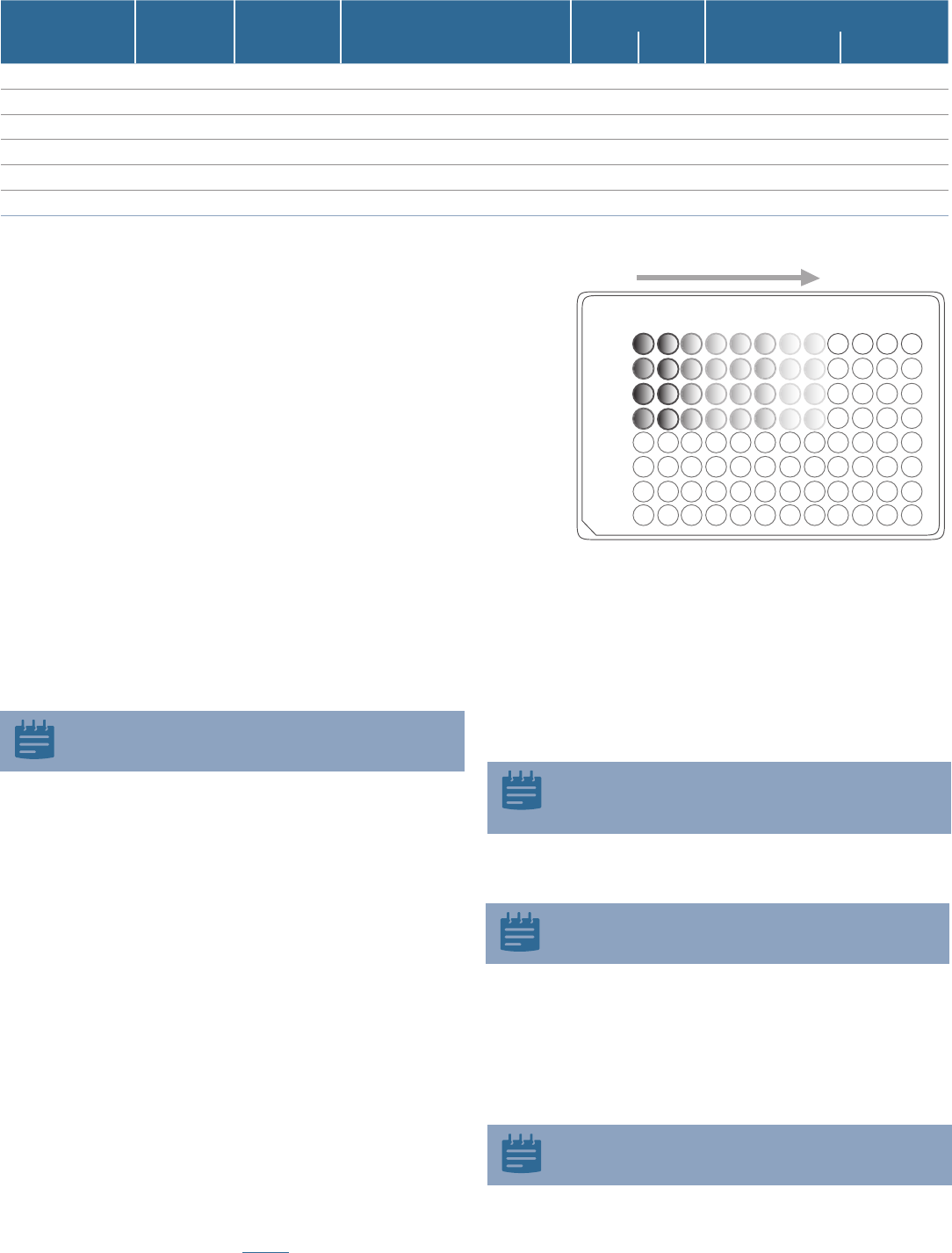
2. Make dilutions of the viral stock in a round bottom 96-well plate
using serum-free media. Utilize the plate as shown in (Figure 6)
using one row for each virus stock to be tested. Use the procedure below
(starting at step 4) for dilution of the viral stocks. The goal is to produce a
series of 5-fold dilutions to reach a final dilution of 390,625-fold.
3. Add 80 µL of serum-free media to each well.
4. Add 20 µL of thawed virus stock to each corresponding well in
column 1 (five-fold dilution).
If you have generated a lentiviral stock of the expression
control (such as GIPZ non-silencing construct), we
recommend titering this stock as well.
The following day, each well should be no more
than 40-50% confluent.
Pipette contents of well up and down 10-15 times. Discard
pipette tip.
Figure 6. Five-fold serial dilutions of virus stock.
Virus stock 1
Virus stock 2
Virus stock 3
Virus stock 4
Dilution Plate
A
1 2345678910 11 12
B
C
D
E
F
G
H
horizondiscovery.com
Protocol V – Puromycin selection
Determining antibiotic dose response (Kill curve)
In order to generate stable cell lines expressing the transgene of interest, it
is important to determine the minimum amount of antibiotic required to kill
non-transfected cells. A simple procedure to test this is as follows:
1. Day 1: Using the same cell type and relative cell densities to be used in
subsequent transfection or transduction procedures, plate cells
and culture overnight.
2. Day 2: Replace complete growth medium with growth medium
supplemented with a range of puromycin concentrations (0-15 μg/mL),
including untreated control cells with no antibiotic added.
3. Day 4: Refresh medium and assess viability.
4. Replace medium with fresh medium supplemented with the appropriate
concentration of puromycin every 2-3 days depending on the growth
of cells.
5. Examine cells daily and identify the minimal concentration of antibiotic
that efficiently kills all non-transfected/transduced cells between
3-6 days following addition of puromycin.
Puromycin selection
If adding antibiotic for selection, use the appropriate concentration as
determined based on the above kill curve.
1. Add medium containing antibiotic 24 or hours post-transfection or
post-transduction, respectively, to begin selection
2. Cells can be harvested for transgene expression 24-72 hours after
starting selection.
3. If longer selection is required for cells to be confluent, replace selective
medium approximately every 2-3 days.
4. Monitor the cells daily and observe the percentage of surviving cells.
Cells surviving selection will be expressing the transgene.
5. If generating stable cell lines (optional), select and grow for 10-15 days.
6. Once non-transfected cells are eliminated and/or you have selected for
stably transfected cell lines if desired, you can proceed to assay for target
gene expression. RT-qPCR, Western blot analysis or other appropriate
functional assay can be used; compare treated samples to untreated,
reporter alone, non-silencing control, or other controls as appropriate.
Protocol VI – Packaging lentivirus
The pGIPZ vector is tat dependant, so you must use a packaging system
that expresses the tat gene. For packaging our lentiviral shRNA constructs,
we recommend the Trans-Lentiviral shRNA Packaging Kit (Cat #TLP5912
or TLP5917). The Trans-Lentiviral shRNA Packaging System allows creation
of a replication-incompetent, HIV-1-based lentivirus which can be used to
deliver and express your shRNA of interest in either dividing or non-dividing
mammalian cells. The Trans-Lentiviral shRNA Packaging System uses a
replication-incompetent lentivirus based on the trans-lentiviral system
developed by Kappes (Kappes 2001). For protocols and information on
packaging pGIPZ with our Trans-Lentiviral shRNA Packaging System, please
see the product manual available on our website.
It is important to wait at least 24 hours after transfection
before beginning selection.

Once you have generated a lentiviral stock with a suitable titer, you are ready to
transduce the lentiviral vector into the mammalian cell line of choice and assay
for gene silencing
Multiplicity of infection (MOI)
To obtain optimal silencing of your shRNA, you will need to transduce the
lentiviral vector into your mammalian cell line of choice using a suitable MOI.
MOI is defined as the number of transducing units per cell.
Determining the optimal MOI
A number of factors can influence determination of an optimal MOI including
the nature of your mammalian cell (actively dividing versus non-dividing), its
transduction efficiency, your application of interest, and the nature of your gene
of interest. If you are transducing your lentiviral construct into the mammalian cell
line of choice for the first time, after you have titered the lentiviral particles, we
recommend using a range of MOIs (for example, 0, 0.5, 1, 2, 5, 10, 20) to determine
the MOI required to obtain optimal expression for your particular application.
It should be noted that to achieve single copy knockdown, an MOI of 0.3 is
generally used, as less than 4% of your cells will have more than one insert.
Protocol VIII – Transduction
Transduction of target cells
The protocol below is optimized for transduction of the lentiviral particles into
HEK293T, OVCAR-8 or MCF7 cells in a 24-well plate using serum-free media. If a
different culture dish is used, adjust the number of cells, volumes, and reagent
quantities in proportion to the change in surface area (Table 7). It is strongly
recommended that you optimize transduction conditions to suit your target cell
line to provide for the highest transduction efficiency possible.
It is preferable that transduction be carried out in medium that is serum
free and antibiotic free. A reduction in transduction efficiency occurs in the
presence of serum; however it is possible to carry out successful transductions
with serum present; you will have to optimize the protocol according to your
cells of interest.
5. With new pipette tips, transfer 20 µL from each well of column 1
to the corresponding well in column 2.
6. With new pipette tips, transfer 20 µL from each well of column 2 to the
corresponding well in column 3.
7. Repeat transfers of 20 µL from columns 3 through 8, pipetting up and
down 10-15 times and changing pipette tips between each dilution.
8. Label 24-well plate as shown in (Figure 7) using one row for each virus
stock to be tested.
9. Remove culture media from the cells in the 24-well plate.
10. Add 225 µL of serum-free media to each well.
11. Transduce cells by adding 25 µL of diluted virus from the original
96-well plate (Figure 6) to a well on the 24-well destination plate
(Figure 7) containing the cells. For example, transfer 25 µL from well
A2 of the 96-well plate into well A1 in the 24-well plate (Table 6).
12. Incubate transduced cultures at 37 °C for 4 hours.
13. Remove transduction mix from cultures and add 1 mL of DMEM
(10% FBS, 1% Pen-Strep).
14. Culture cells for 48 hours.
15. Count the TurboGFP expressing cells or colonies of cells (Figure 8).
16. Transducing units per mL (TU/mL) can be determined using the
following formula:
# of TurboGFP positive colonies counted × dilution factor × 40 = # TU/mL:
Example: 55 TurboGFP positive colonies counted in well A3
55 (TurboGFP positive colonies) × 625 (dilution factor) × 40 = 1.38 ×
10
6
TU/mL
Pipette up and down 10-15 times and discard pipette tip.
Pipette up and down 10-15 times and discard pipette tip.
It is strongly recommended that you use a high quality
multichannel pipettor when performing multiple dilutions.
Pre-incubate the dilutions of the virus stock for 5 minutes
at room temperature.
Count each multi-cell colony as 1 transduced cell, as the
cells will be dividing over the 48 hour culture period. Figure
8 illustrates this principle of cell counting.
A
B
C
D
1 2 3 4 5 6
Virus stock 1
Virus stock 2
Virus stock 3
Virus stock 4
Figure 7. Twenty four well tissue culture plate, seeded with HEK293T cells, used
to titer the virus.
Figure 8. Examples of individual colonies.
A B
Well (Row A, B, C, or D)
Volume diluted
virus used
Dilution factor
Originating
(96-well plate)
Destination
(24-well plate)
A1 25 μL 5 *
A2 A1 25 μL 25
A3 A2 25 μL 125
A4 A3 25 μL 625
A5 A4 25 μL 3125
A6 A5 25 μL 15625
A7 A6 25 μL 78125
A8 25 μL 390625 *
Table 6. Example of set up for dilutions.
*Please note that when expecting very high or very low titers, it would be advisable to
include either well 8 or well 1 respectively.
horizondiscovery.com

1. On day 0, plate 5 × 10
4
cells per well in a 24-well plate. Incubate
overnight. You will be using full growth medium (with serum) at
this stage.
2. The next day (day 1), remove the medium and add an appropriate
amount of the virus to acheive the MOI you wish to use. Set up
all desired experiments and controls in a similar fashion. Bring
the total volume of liquid up so that it just covers the cells
efficiently with serum-free media (see Table 7 for guidelines).
If you are using concentrated virus you are likely to use a small
volume of virus if you are using unconcentrated virus, you will
find you need more virus volume.
3. Approximately 4-6 hours post-transduction, add an additional 1 mL
of full medium (serum plus Pen-Strep if you are using it) to your cells
and incubate overnight.
At 48 hours post-transduction examine the cells microscopically for
the presence of reporter gene (TurboGFP) expression as this will be
your first indication as to the efficiency of your transduction.
a. If adding puromycin, use the appropriate concentration as
determined based on the kill curve (see Protocol V). Incubate cells
with the selection medium.
b. Approximately every 2-3 days replace with freshly prepared
selective medium.
c. Monitor the cells daily and observe the percentage of surviving
cells. Once the non-transduced control cells are dead, the surviving
cells in the transduced wells will be expressing the shRNA
Optimum effectiveness of the puromycin selection should be
reached in 4-6 days with puromycin dependent upon the
concentration of puromycin chosen from the kill curve.
We have experienced low toxicity with transduction in the
cell lines tested, therefore removal of virus is not required
for many cell lines. In our experience, higher transduction
efficiencies have been achieved if the virus is not removed
after 6 hours. However, if toxicity is a problem, aspirate
the mixture after 4-6 hours and replace with fresh growth
medium. Additionally, fresh growth medium should be
replenished as required for continued cell growth.
The higher the MOI you have chosen, the more copies
of the shRNA and puromycin resistance gene you will
have per cell. When selecting with puromycin, it is worth
remembering that at higher MOIs, cells containing multiple
copies of the resistance gene can withstand higher
puromycin concentrations than those at lower MOIs. Adjust
the concentration of puromycin to a level that will select
for the population of transduced cells you require for your
application without going below the minimum antibiotic
concentration you have established in your kill curve.
Once your transduction efficiency is at an acceptable level (with or without
puromycin selection performed post-transduction), you can proceed to assay
cells for reduction in gene expression or fluorescent reporter activity by reverse
transcription quantitative/real-time PCR (RT-qPCR), Western blot analysis or
other appropriate functional assay. Compare target gene to untreated, reporter
alone (empty vector), non-silencing shRNA, or other negative controls.
Protocol IX – Determining relative
transduction eciency
Follow the procedure below to determine the relative transduction efficiency
of purchased GIPZ lentiviral particles (Cat #VGH5518, VGM5520, VGH5526). This
protocol should only be used with purchased GIPZ shRNA individual clones in
viral particle format.
Prior to transducing with purchased GIPZ shRNA individual clones in viral
particle format, we recommend determining the relative transduction efficiency
of your cell type. Lentiviral titers provided with purchased GIPZ lentiviral
particles have been calculated by transducing HEK293T cells. Transduction
efficiencies vary significantly by cell type.
The relative transduction efficiency of your cells may be estimated by
determining the functional titer of a control virus such as GIPZ Non-silencing
control viral particles (Cat #RHS4348) in your cells of interest.
Follow the procedure below to determine the functional titer of the GIPZ
Non-silencing control shRNA viral stock in the mammalian cell line of your
choice. The following conditions have been optimized for transduction of
HEK293T cells. When determining the relative transduction efficiency of
your cell type, use the transduction conditions that have been optimized
for your cells of interest.
1. The day before transduction (day 0), seed a 24-well tissue culture plate
with your cells at 5 × 10
4
cells per well in their respective medium.
2. Make dilutions of the Non-silencing control shRNA viral stock in a round
bottom 96-well plate using serum-free medium. Utilize the plate as shown
in Figure 9 with one row for each replicate (we recommend performing
at least two replicates). Use the procedure below for dilution of the viral
stock. The goal is to produce a series of five-fold dilutions to reach a final
dilution of 390, 625-fold.
Optimal length of incubation from the start of transfection
to analysis is dependent on cell type, gene of interest,
and the stability of the mRNA and/or protein being
analyzed. RT-qPCR generally gives the best indication of
mRNA expression and gene silencing. The use of Western
blot analysis to determine knockdown is dependent on
quantity and quality of the protein, its half-life, and the
sensitivity of the antibody and detection systems used.
The following day, each well should be no more than
40-50% confluent.
When visualizing TurboGFP expression, if less than
90% of all cells are green, it is recommended in these
cases to utilize puromycin selection in order to reduce
background expression of your gene of interest from
untransduced cells.
horizondiscovery.com
Tissue
culture dish
Surface area
per well (cm
2
)
Suggested total serum-free medium
volume per well (mL)
100 mm 56.0 5.0
60 mm 20.0 2.0
35 mm 8.0 1.0
6-well 9.4 1.0
12-well 3.8 0.5
24-well 1.9 0.25
96-well 0.3 0.1
Ta ble 7. Suggested volumes of media per surface area per well of adherent cells.

3. Add 40 uL of serum-free media to each well in column.
4. Add 80 uL of serum-free media to each well of columns 2-8.
5. Add 10 µL of thawed control shRNA virus stock to each well in column 1
(five-fold dilution).
6. With new pipette tips, transfer 20 µL from each well of column 1 to the
corresponding well in column 2.
Pipette up and down 10-15 times and discard pipette tip.
7. With new pipette tips, transfer 20 µL from each well of column 2 to the
corresponding well in column 3.
Pipette up and down 10-15 times and discard pipette tip.
8. Repeat transfers of 20 µL from columns 3 through 8, pipetting up and
down 10-15 times and changing pipette tips between each dilution.
It is strongly recommended that you use a high-quality
multichannel pipettor when performing multiple dilutions.
Incubate the dilutions of the virus stock for 5 minutes at room
temperature.
9. Label a 24-well plate as shown in Figure 10 using one row for each
replicate.
10. Remove culture medium from the cells in the 24-well plate.
11. Add 225 µL of serum-free medium to each well.
12. Transduce cells by adding 25 µL of diluted control shRNA lentivirus
from the original 96-well plate (Figure 9) to a well on the 24-well
destination plate (Figure 10) containing the cells. For example,
transfer 25 µL from well A2 of the 96-well plate into well A1in the
24-well plate (Table 8).
13. Incubate transduced cultures at 37 °C for 4-6 hours.
If desired, include 8 μg/mL polybrene in the
dilution medium.
Pipette contents of well up and down 10-15 times.
Discard pipette tip.
Post dox 72 hours
14. Add 1 mL of your medium (normal serum concentration).
15. Culture cells for 72 hours.
16. Count the TurboGFP expressing cells or colonies of cells (Figure 11). Count
each multi-cell colony as 1 transduced cell, as the cells will be dividing
over the 72 hour culture period. Figure 11 illustrates this principle of
cell counting. Count the number of TurboGFP expressing colonies in
wells corresponding to at least two viral dilutions.
17. Transducing units per ml (TU/mL) can be determined using the following
formula: # of TurboGFP positive colonies counted × dilution factor × 40 =
# TU/mL
Example: 55 TurboGFP positive colonies counted in well A3 55 (TurboGFP
positive colonies) × 625 (dilution factor) × 40 =
1.38 × 10
6
TU/mL.
18. The functional titer calculated for your cell line under your experimental
conditions can be used to determine the relative transduction efficiency
of your cell type by using the following formula: Functional titer of Non-
silencing control shRNA virus stock in your cell line ÷ Titer of Non-silencing
control shRNA virus stock as calculated in HEK293T = Relative transduction
efficiency.
For example, if the titer of the Non-silencing control shRNA virus stock in
HEK293T (as provided on the product specification sheet) is 6.9 × 10
6
TU/mL
and the functional titer of the control shRNA virus stock in your cell line is 1.38
× 10
6
TU/mL, the relative transduction efficiency of your cell type is 0.2. To
extrapolate the average functional titer of the provided GIPZ viral particles,
multiply the average titer of each plate as provided on the product specification
sheet by the relative transduction efficiency of your cell type. In our example,
if the titer of the GIPZ viral particles in HEK293T cells is 2 × 10
6
TU/mL and the
relative transduction efficiency of your cell type is 0.2, the extrapolated average
functional titer of that plate your cell type is 4 × 10
5
TU/mL.
25 μLof diluted virus was added to the cells. This is
1/40th of a mL.
A
B
C
D
1 2 3 4 5 6
Virus stock 1
Virus stock 2
Virus stock 3
Virus stock 4
Figure 10. Twenty-four well tissue culture plate, seeded
with HEK293T cells, used totiter the virus.
horizondiscovery.com
Figure 9. Five-fold serial dilutions of virus stock.
Virus stock 1
Virus stock 2
Virus stock 3
Virus stock 4
Dilution Plate
A
1 2345678910 11 12
B
C
D
E
F
G
H
Figure 11. Examples of individual colonies.
A B
*Please note that when expecting very high or very low titers, it would be
advisable to include either well 8 or well 1 respectively.
Well (row A, B, C, or D)
Volume diluted
virus used
Dilution
factor
Originating
(96-well plate)
Destination
(24-well plate)
A1 25 μL 5 *
A2 A1 25 μL 25
A3 A2 25 μL 125
A4 A3 25 μL 625
A5 A4 25 μL 3125
A6 A5 25 μL 15625
A7 A6 25 μL 78125
A8 25 μL 390625 *
Table 8. Example of set up for dilutions.

Once the relative transduction efficiency of the GIPZ virus has been
established in your cell line, use the optimized transduction conditions
determined in Protocol VIII to transduce your cell line with the purchased
GIPZ shRNA individual clones in viral particle format. If the titer of the non-
silencing control shRNA virus is not satisfactory in your cell line you might
consider choosing a different cell line more permissive to transduction by
lentivirus before proceeding.
Protocol X – PCR
QPCR experimental recommendations
One of the biggest challenges of any qPCR experiment is to obtain
reproducible and reliable data. Due to the sensitivity of this multi-step
technique, care must be taken to ensure results obtained are accurate and
trustworthy (see Bustin et al., 2009).
1. Experimental samples should be run at least in duplicate. It should
be noted that with duplicate experiments it will not be possible to
assign error bars to indicate consistency from experimental sample
to experimental sample. Using triplicate samples or higher will
enable error bars to be assigned indicating the level of experimental
variation.
2. Reverse Transcriptase reactions for cDNA synthesis should always
include a No Template Control (NTC) and No Reverse Transcriptase
(no RT) control to check for reagents contamination and the
presence of contaminating DNA, respectively. Use a robust reverse
transcriptase enzyme for cDNA synthesis such as the Thermo
Scientific
™
Maxima
™
cDNA Synthesis Kit for RT-qPCR (Cat #K1641).
3. We have found that normalizing the RNA concentration prior to
cDNA synthesis will increase consistency downstream.
4. qPCR should be done at least in triplicate. Again, it should be noted
that with duplicate reactions it will not be possible to assign error
bars to indicate the consistency in your qPCR reactions. Using
triplicate samples or higher will enable error bars to be assigned
indicating the level of variation between qPCR reactions. Use
validated primer sets for SYBR-based assays or primers/probe for
probe-based assays.
5. Make sure the mRNA you are using as your internal reference control
for qPCR is expressed at a level higher than your target gene's
message.
6. Use only high-quality calibrated pipettes, in conjunction with well
fitting barrier tips.
7. When pipetting, take the time to visually inspect the fluid in the
pipette tip(s) for accuracy and lack of bubbles, especially when using
a multi-channel pipette.
8. Be sure to spin your qPCR plate prior to loading in the real-time
instrument in order to collect the sample at the bottom of the well
and eliminate any bubbles that may have developed.
9. With regard to knockdown experiments using shRNA, it is vitally
important that you greatly reduce if not eliminate entirely those
cells which are not transduced or transfected from the population.
This can be done in several ways: increase the efficiency of your
transfection, use a higher mulitplicity of infection (MOI) for
your transduction, utilize the puromycin selection marker and
select against those cells that do not contain the shRNA or utilize
fluorescent sorting to select against those cells that do not contain
the shRNA.
10. Always utilize the non-silencing control as a reference for target gene
expression, as opposed to an untreated sample. The non-silencing
treated samples will most accurately reproduce the conditions in your
experimental samples. The non-silencing best controls for changes in
qPCR internal reference gene expression.
11. You may also use an untreated sample to indicate substantial
changes in target gene expression as seen in the non-silencing control
due to generic consequences of viral transduction and transgene
expression. However, it should be noted that small changes in
expression levels between an untreated sample and the non-silencing
control are to be expected.
12. Cq values greater than 35 should be avoided as they tend to be
more
variable. Samples with such high Cq values should be repeated at
higher cDNA concentrations and with a lower expressing qPCR internal
reference control (such as TBP).
13. Cq values less than 11 for the qPCR internal reference control should be
avoided as it is difficult to determine a proper background subtraction
using these values. If this occurs, use Cq values from both your internal
reference control as well as your experimental target to determine an
optimum cDNA concentration.
14. It may be necessary to change internal reference controls if conditions
in steps 12 and 13 cannot be simultaneously met.
100%
113%
88%
93%
120%
140%
Control GAPDH EG5 PP1B
100%
80%
60%
40%
20%
0%
Residual Gene
Activity
Figure 12. Non-silencing lentiviral shRNA control does not knockdown common
endogenous genes. The above data represents the baseline amount of GAPDH, EG5 or
PP1B mRNA set at 100% in the control. The relative amounts of each of these mRNAs are
then represented after treatment with non-silencing shRNA of these genes.
Figure 13. HEK293T cells were transduced with lentiviral particles expressing GAPDH
or Non-silencing shRNA at variable MOIs ranging from 9-48. The graph depicts the
residual levels of GAPDH relative to Non-silencing control.
120%
Untrans-
duced
Control
NS#1 NS#1 NS#1 GAPDH GAPDH GAPDH
100%
80%
60%
40%
20%
0%
12%
28%
34%
100%100%100%100%
Residual GAPDH
Expression
horizondiscovery.com

Figure 15. The characteristic phenotype observed by the targeting of the EG5 (KIF11)
gene results in the formation of half spindles, mitotic arrest and monoastral microtubular
arrays (green, see the cell on the left). By contrast, normal cells show bipolar spindles
and microtubule networks in mitosis and in interphase (see the cell on the right). The
comparative expression of EG5 (red) between the cell on the left and the right shows
the extensive knockdown of EG5 in the cell displaying the phenotype (left). The cells
were visualized at 100x magnification using aLeica DMIRB fluorescence microscope.
HEK293T cells were stained for tubulin (anti-tubulin, green), DNA (DAPI, blue) and EG5
(anti-EG5, red).
OVERLAY ANTI-TUBULIN
ANTI-EG5 DAPI
Related Reagents Dharmacon Cat #
GAPDH Verified Positive Control* RHS4371
EG5 Verified Positive control* RHS4480
Non-silencing Verified Negative Control* RHS4346
DharmaFECT kb transfection reagent 1 mL 2006-01
GIPZ shRNA Empty Vector RHS4349
Trans-Lentiviral shRNA Packaging System TLP5912
Trans-Lentiviral shRNA Packaging System with HEK293T Cells TLP5917
Table 9. Related Reagents.
*These items also available in the GIPZ lentiviral transfection starter kit (Cat #RHS11851).
Controls and validation
RNAintro shRNA starter kits
The use of vector-based RNAi for gene silencing is a powerful and versatile tool.
Successful gene silencing in vitro is dependent on several variables including 1)
The target cell line being studied, 2) transfection and transduction efficiency,
3) abundance of the mRNA or protein of interest in the target cell line, 4) half
life of the protein, and 5) robust experimental protocols. For all these reasons,
it is important to run controlled experiments where the transfection and
transduction efficiencies are as high as possible and measurable.
Controls are a critical part of a gene silencing experiment. They enable
accurate representation of knockdown data and provide confidence in the
specificity of the response. Changes in the mRNA or protein levels in cells
treated with negative or non-silencing controls reflect non-specific responses
in cells and can be used as a baseline against which specific knockdown
can be measured. Positive controls are useful to demonstrate that your
experimental system is functional.
Controls
The EG5 and GAPDH GIPZ lentiviral shRNA vectors have been validated as
positive controls for RNAi experiments performed using the GIPZ shRNA-
containing lentiviral vectors. These shRNA have been tested in transduction
based experiments and have shown efficient knockdown at both mRNA and
protein levels. The EG5 control has been validated to knockdown human EG5 by
RT-qPCR (Figure 14 and 15) and in situ hybridization of cells in tissue culture. The
GAPDH control has been validated to knockdown human and mouse GAPDH
by RT-qPCR (Figure 13). The GIPZ Non-silencing lentiviral shRNA vector has been
validated as a negative control for RNAi experiments performed using the GIPZ
shRNA-containing lentiviral vectors (Figure 12).
Frequently Asked Questions (FAQs)
What clones are part of my library collection?
A USB containing the data for this collection will be shipped with each
collection. This file contains the location and accession number for each
construct in the collection.
Where can I find the sequence of an individual shRNA construct?
If you are looking for the sequence of an individual shRNA construct, you can
search for the clone on our website (horizondiscovery.com). Enter the catalog
number or clone ID of your construct into the search at the top of the page. You
should see your product in the catalog number section of the results. Click on
the plus sign to expand the details for this clone and select the Sequence tab.
Which antibiotic should I use?
You should grow all GIPZ shRNA constructs in 2x LB broth medium with both
25 μg/mL zeocin and 100 μg/mL carbenicillin for archive replication. For
plasmid preparations, grow the constructs in 2x LB broth medium containing
only 100 ug/mL carbenicillin.
What packaging cell line should I use for making lentivirus?
The GIPZ shRNA vector is tat dependant, so a packaging system that
expresses the tat gene. For packaging our lentiviral shRNA constructs, we
recommend the Trans-Lentiviral shRNA Packaging Kit (TLP5912, TLP5917). The
Trans-Lentiviral Packaging Kit allows creation of a replication-incompetent
(Shimada, et al. 1995), HIV-1-based lentivirus which can be used to deliver
and express your gene or shRNA of interest in either dividing or non-dividing
mammalian cells. The Trans-Lentiviral Packaging Kit uses a replication-
incompetent lentivirus based on the trans-lentiviral system developed
by Kappes (Kappes and Wu et al. 2001). For protocols and information on
packaging GIPZ shRNA with our Trans-Lentiviral shRNA Packaging Kit, please
see the product manual available at here.
Can I use any 2
nd
generation packaging system with the pGIPZ vector?
The pGIPZ vector is tat dependant, so you must use a packaging system that
expresses the tat gene.
What does the number 40 refer to in the formula for the calculation of titer?
The titer units are given in transducing units (TU) per mL, so the number 40 is
used to convert the 25 µL used in the titration (“volume of diluted virus used”,
Table 6) to one milliliter.
What is the sequencing primer for pGIPZ?
The pGIPZ sequencing primer is 5' - GCATTAAAGCAGCGTATC - 3'
Note: The binding site lies from base 5820-5842 and runs in the reverse
complement direction. The melting temperature of this 18mer = 52.7 °C.
Where do you purchase puromycin?
We purchase puromycin from Fisher Scientific Cellgro (Cat #BP2956-100).
How many transfections are available in each volume size of
DharmaFECT kb transfection reagent?
The number of transfections that can be performed depends on the size of the
culture dish used.
120%
NS#1
MOI 3.5
NS#1
MOI 8.5
NS#1
MOI 17
EG5
MOI 3.5
EG5
MOI 8.5
EG5
MOI 17
100%
80%
60%
40%
20%
0%
17%
18%
29%
100%100%100%
Residual EG5
Expression
Figure 14. HEK293T cells were transduced with lentiviral particles expressing EG5 or
non-silencing shRNA at MOIs of 3.5, 8.5 and 17. The graph depicts the residual levels of
EG5 relative to its non-silencing control.

Troubleshooting
For help with transfection or transduction of your lentiviral constructs, please
email technical support at technical@horizondiscovery.com with the answers
to the questions below, your sales order or purchase order number and the
catalog number or clone ID of the construct with which you are having trouble.
1. Are you using direct transfection or transduction into your cell line?
2. What was the 260/280 ratio of DNA? Over 1.8?
3. What was the transfection efficiency if you used direct transfection?
What transfection reagent was used?
4. Were positive and negative knockdown controls used (such as our
GAPDH or EG5 validated positive controls and the validated non-
silencing negative control)?
5. What were the results of the controlled experiments?
6. How was knockdown measured (for example real-time RT-qPCR or
western blot analysis)?
7. What is the abundance and the half-life of the protein? Does the protein
have many isoforms?
8. What packaging cell line was used if you are using transduction rather
than transfection?
9. What was your viral titer?
10. What was your MOI?
11. Did you maintain the cells in puromycin selection media after
transfection or transduction?
12. How much time elapsed from transfection/transduction to
puromycin selection?
If transfection into your cell line is unsuccessful, you may need to
consider the following list of factors influencing successful transfection.
1. Concentration and purity of plasmid DNA and nucleic acids—determine
the concentration of your DNA using 260 nm absorbance. Avoid
cytotoxic effects by using pure preparations of nucleic acids.
2. Insufficient mixing of transfection reagent or transfection complexes.
3. Presence of antibiotics in transfection medium—the presence of antibiotics
do not interfere with both DNA/DharmaFECT kb complex formation and cell
transfection. This is not the case for other transfection reagents.
4. Cell history, density, and passage number—it is very important to use
healthy cells that are regularly passaged and in growth phase. The highest
transfection efficiencies are achieved if cells are plated the day before;
however, adequate time should be given to allow the cells to recover from
the passaging (generally > 12 hours). Plate cells at a consistent density to
minimize experimental variation. If transfection efficiencies are low or
reduction occurs over time, thawing a new batch of cells or using cells with
a lower passage number may improve the results.
If transduction into your cell line is unsuccessful, you may need to
consider the following list of factors influencing successful transduction.
1. Transduction efficiency is integrally related to the quality and the
quantity of the virus you have produced. Factors to consider when
transducing include MOI (related to accurate titer in the target cell line),
the presence of serum in the media, the use of polybrene in the media,
length of exposure to virus, and viral toxicity to your particular cells.
2. High quality transfer vector DNA and the appropriate and efficient viral
packaging are required to make high quality virus able to transduce
cells effectively.
3. See also suggestions 3–5 for factors influencing successful
transfection (above).
References
Cited References and additional suggested reading
1. Bartel, D. P. (2004). microRNAs: genomics, biogenesis, mechanism, and
function. Cell 116(2): 281- 97.
2. Boden, D., O. Pusch, et al. (2004). Enhanced gene silencing of HIV-1 specific
siRNA using microRNA designed hairpins Nucleic Acids Res 32(3): 115 4 -8.
3. Chendrimada, T. P., R. I. Gregory, et al. (2005). TRBP recruits the Dicer
complex to Ago2 for microRNA processing and gene silencing. Nature
436(7051): 740-4.
4. Cleary, M. A., K. Kilian, et al. (2004). Production of complex nucleic acid
libraries using highly parallel in situ oligonucleotide synthesis.
Nat Methods 1(3): 241-8.
5. Cullen, B. R. (2004). Transcription and processing of human microRNA
precursors. Mol Cell 16(6): 861-5.
6. Cullen, B. R. (2005). RNAi the natural way. Nat Genet 37(11): 1163-5.
7. Dickins, R. A., M. T. Hemann, et al. (2005). Probing tumor phenotypes using
stable and regulated synthetic microRNA precursors. Nat Genet 37(11):
1289 -95.
8. Editors of Nature Cell Biology (2003). Whither RNAi? Nat Cell Biol 5(6):
489-90.
9. Elbashir, S. M., J. Harborth, et al. (2001). Duplexes of 21-nucleotide
RNAs mediate RNA interference in cultured mammalian cells. Nature
411(6836): 494-8.
10. Fire, A., S. Xu, et al. (1998). Potent and specific genetic interference by
double-stranded RNA in Caenorhabditis elegans. Nature 391(6669):
806 -11.
11. Gregory, R. I., T. P. Chendrimada, et al. (2005). Human RISC couples
microRNA biogenesis and posttranscriptional gene silencing.
Cell 123(4): 631-40
12. Kappes, J. C. and X. Wu (2001). Safety considerations in vector
development. Somat Cell Mol Genet 26(1-6):147-58.
13. Kappes, J. C., X. Wu, et al. (2003). Production of trans-lentiviral vector
with predictable safety. Methods Mol Med 76: 449-65.
14. Paddison, P. J., J. M. Silva, et al. (2004). A resource for large-scale
RNA-interference-based screens in mammals. Nature 428(6981): 427-31.
15. Shimada, T., et.al. (1995). Development of vectors utilized for gene
therapy for AIDS. AIDS 4.
16. Silva, J. M., M. Z. Li, et al. (2005). Second-generation shRNA libraries
covering the mouse and human genomes. Nat Genet 37(11): 1281-8.
Lable licenses
The shRNA Products, use and applications, are covered by pending and
issued patents. Certain Label licenses govern the use of the products, these can
be found at dharmacon-licensing-statements. It is each Buyer’s responsibility to
determine which intellectual property rights held by third parties may restrict
the use of Products for a particular application. Please review the Label Licenses
governing all use of the shRNA and gene expression products.
To find the contact information in your country for your technology of interest, please visit us at
horizondiscovery.com/contact-us
Horizon Discovery, 8100 Cambridge Research Park, Waterbeach, Cambridge, CB25 9TL, United Kingdom
©2020 The Horizon logo and other trademarks are the property of Horizon Discovery Limited, unless otherwise stated. DHARMACON, GIPZ and DHARMAFECT are trademarks of
Dharmacon Inc. TURBOGFP is a trademark of Evrogen JS . HXB2 is a trademark of GenBank. ZEOCIN, FASTDIGEST, GENEJET, and MAXIMA are trademarks of Thermo Fisher Scientific, Inc
For more information
V101120
